 Insects have social lifestyles that are often organized in castes. Within the insect community, different individuals specialize, each having a unique role. This efficient method of doling out the workload, ultimately, is believed to be why social insect lifestyles are successful. However, how it’s determined who does what is really pretty cool.
Insects have social lifestyles that are often organized in castes. Within the insect community, different individuals specialize, each having a unique role. This efficient method of doling out the workload, ultimately, is believed to be why social insect lifestyles are successful. However, how it’s determined who does what is really pretty cool.
You’re probably familiar with the system used by the honeybee (Apis mellifera), where one queen focuses on producing more honey making babies while sterile worker bees support her. This globally essential insect has gotten a lot of attention lately because populations have diminished worldwide, as has been pointed out in a suite of articles here.
In honeybees, bees with different jobs are genetically the same. In fact, three different types (phenotypes) of honeybees are produced from the same genome. Fertilized embryos (diploids) can become functionally sterile female worker OR the reproductive queen, while unfertilized eggs (haploids) produce male drones. For every queen, there are up to 50,000 workers keeping things running smoothly. But if they’re essentially genetically identical, what makes these guys different?
 Turns out, it’s nurture (not nature) that’s driving things along. The queen and the workers receive different types and amounts of food (royal jelly only for queenie), as well as more luxuriously large cells to develop in, all of which turns on different developmental pathways. The amount of space that a larva develops in changes levels of DNA methylation. Basically, this is a way switching the DNA “off” or “on” for certain genes, without actually changing the DNA.
Turns out, it’s nurture (not nature) that’s driving things along. The queen and the workers receive different types and amounts of food (royal jelly only for queenie), as well as more luxuriously large cells to develop in, all of which turns on different developmental pathways. The amount of space that a larva develops in changes levels of DNA methylation. Basically, this is a way switching the DNA “off” or “on” for certain genes, without actually changing the DNA.
The study of these changes in gene expression is called epigenetics, and it turns out, it’s pretty dang important. Why might we care about this system? Well, apart from that it’s really interesting, commercial beekeepers rear their own queens often from worker bee larvae, and considering the latest observations of honeybee colony decline, the queen quality might have a role in exacerbating failure rates.
Xu Jiang He and colleagues recently published an article in Molecular Ecology analyzing genomic regulation in both worker and queen bees. They explored how different variables in queen rearing affect queen quality. Their findings suggest that the environment a queen is raised in can lead to a less than perfect queen, proposing that rearing in suboptimal environments could be contributing to global colony failure.
“We found that the domestic rearing practice altered queen morphology and induced epigenetic changes in developing queens….”
 To explore the role of epigenetics in queen rearing, the authors studied three European honeybee colonies. The queens of each colony laid eggs, which were then transplanted for new queen rearing and compared to worker larvae that were transplanted. Specifically, three groups were tested: 1, 2, or 3 day old larvae, labeled QWL1, QWL2, and QWL3, respectively, which were transplanted to queen cells for rearing. After 3 days, the RNA and DNA from 3 larvae of each treatment were collected, they also allowed some to grow to maturity for later measurements. From the mRNA isolated from the total RNA, a cDNA library was put together and sequenced using an Illumina HiSeq 2500 platform. The authors also determined the level of methylation of the DNA extracted.
To explore the role of epigenetics in queen rearing, the authors studied three European honeybee colonies. The queens of each colony laid eggs, which were then transplanted for new queen rearing and compared to worker larvae that were transplanted. Specifically, three groups were tested: 1, 2, or 3 day old larvae, labeled QWL1, QWL2, and QWL3, respectively, which were transplanted to queen cells for rearing. After 3 days, the RNA and DNA from 3 larvae of each treatment were collected, they also allowed some to grow to maturity for later measurements. From the mRNA isolated from the total RNA, a cDNA library was put together and sequenced using an Illumina HiSeq 2500 platform. The authors also determined the level of methylation of the DNA extracted.
The older the worker larvae was before being transplanted to a queen cell, the smaller the final adult queen became. Furthermore, using RNA-Seq, they observed that the number of differentially expressed genes increased with the age of the transplanted worker larva. A bummer for those late transplantations, but turns out that the highest levels of differential expression were in genes involved in immunity, reproductive ability, and longevity, seems important!
“Furthermore, queens from older grafted worker larvae had a higher global DNA methylation level than QWE” (queen eggs that were transplanted to queen cells, e.g. the control)
 This study looked carefully at how the queen development was influenced by both its environment as well as nutrition, understanding how seemingly small variations in queen rearing affect queen quality. Although Xu Jiang He and colleagues found that when older worker larvae are switched to an environment to become a queen, how late the switch is made led to differences in queen longevity. This late transplantation also lead regions of the genome becoming differentially methylated, another a factor that affects the final queen phenotype.
This study looked carefully at how the queen development was influenced by both its environment as well as nutrition, understanding how seemingly small variations in queen rearing affect queen quality. Although Xu Jiang He and colleagues found that when older worker larvae are switched to an environment to become a queen, how late the switch is made led to differences in queen longevity. This late transplantation also lead regions of the genome becoming differentially methylated, another a factor that affects the final queen phenotype.
It turns out that often in commercial practice older worker larvae are preferred for queen rearing since they are apparently hardier and easier to deal with. However, the authors’ findings reveal that it might be worth the extra effort to focus on queen-rearing with younger larvae. Hopefully, studies such as this one, focused on understanding how quality of the queen and therefore the hive might be enhanced will help us tackle continued decreases in bee colonies. The honeybee has also proven to be a cool model system for epigenetics, who would have bee-lieved? (Sorry, I had to get one pun in!)
References
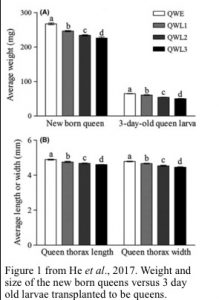
He, X.J., Zhou, L.B., Pan, Q.Z., Barron, A.B., Yan, W.Y. and Zeng, Z.J., 2017. Making a queen: an epigenetic analysis of the robustness of the honeybee (Apis mellifera) queen developmental pathway. Molecular Ecology, 26(6), pp.1598-1607. doi: 10.1111/mec.13990
Ashby, R., Forêt, S., Searle, I. and Maleszka, R., 2016. MicroRNAs in honey bee caste determination. Scientific reports, 6, p.18794. doi:10.1038/srep18794
